Rasmus Fonseca
Data Mining | Structural Biology | Optimization
Projects
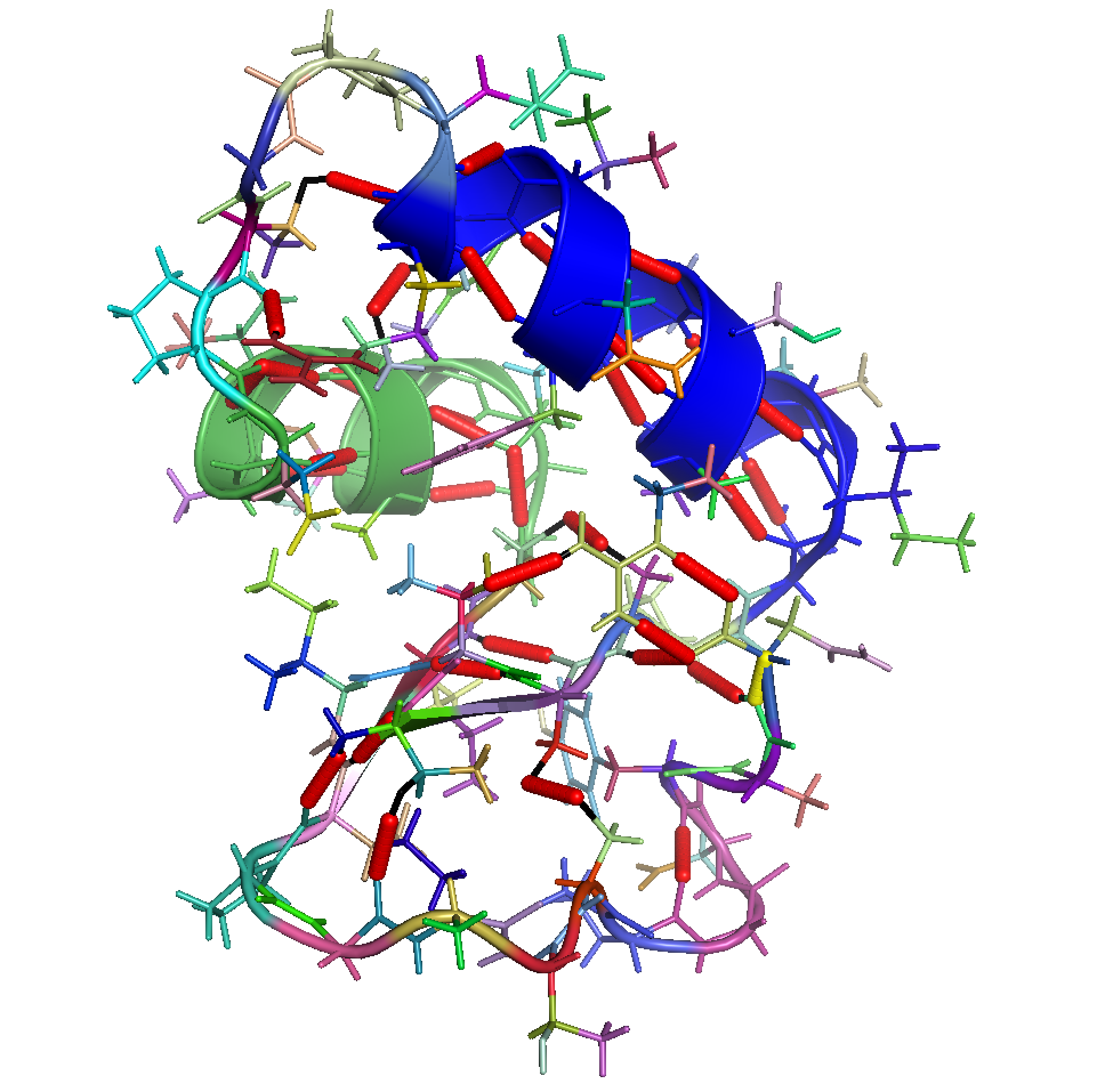
Framework for modeling the conformational flexibility of macromolecules using robotics theory.
GPL3 license, C++

Protein Geometric Algorithm Library. A Java library with algorithms and data structures for analyzing and representing protein structures.
Apache-2.0 license, Java 1.6

Define a regex-like pattern and efficiently search through biological sequences.
GPL2 license, C++
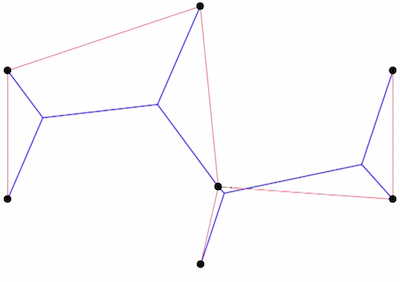
A C++ application for finding optimal Euclidean Steiner trees in any dimension.
GPL2 license, C++
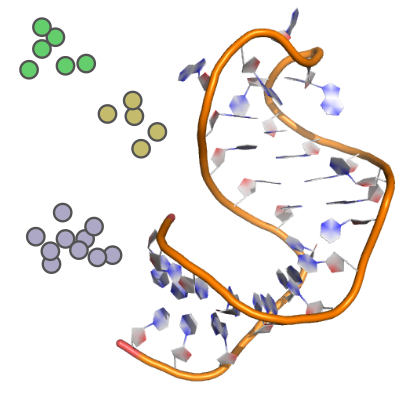
Small C++ application that performs k-medoids and k-centers clustering of protein/RNA/DNA PDB-files using Partitioning Around Medoids (PAM) and the dRMSD metric.
Apache-2.0 license, C++
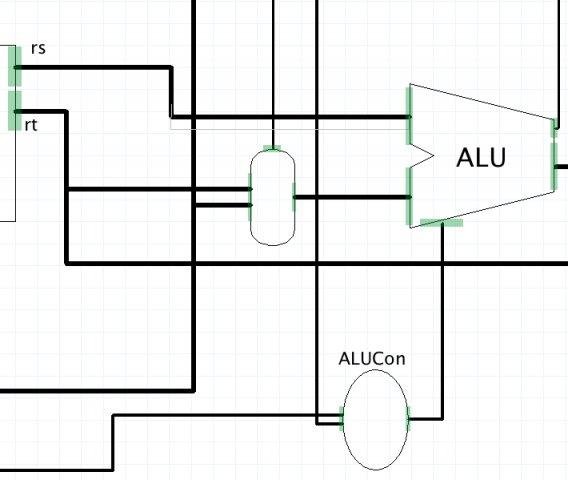
Visual hardware simulator for a machine architecture class. Suitable companion for the book "Computer Organization and Design".
GPL3 license, Java

Library for displaying matrices of real values. Encodes magnitude of entries as circle radius and value as colors.
MIT license, JavaScript
Small library for embedding a slider that allows the user to select an integer range and press a play button.
Apache-2.0 license, JavaScript
Publications
- Uncovering patterns of atomic interactions in static and dynamic structures of proteins A. J. Venkatakrishnan, Rasmus Fonseca, Anthony K. Ma, Scott A. Hollingsworth, Augustine Chemparathy, Daniel Hilger, Albert J. Kooistra, Ramin Ahmari, M. Madan Babu, Brian K. Kobilka, and Ron O. Dror. BioRxiv, 2019
- Conformational transitions of a neurotensin receptor 1–G i1 complex. H E Kato, Y Zhang, H Hu, C-M Suomivuori, F M N Kadji, J Aoki, K K Kumar, R Fonseca, D Hilger, W Huang, N R Latorraca, A Inoue, R O Dror, B K Kobilka, G Skiniotis. Nature, 2019
- Diverse GPCRs exhibit conserved water networks for stabilization and activation. AJ Venkatakrishnan, A K Ma, R Fonseca, N R Latorraca, B Kelly, R M Betz, C Asawa, B K Kobilka, and R O Dror. PNAS, 2019
- Structural insights into the activation of metabotropic glutamate receptors. A Koehl, H Hu, D Feng, B Sun, Y Zhang, M J Robertson, M Chu, T S Kobilka, T Laeremans, J Steyaert, J Tarrasch, S Dutta, R Fonseca, W I Weis, J M Mathiesen, G Skiniotis, and B K Kobilka. Nature, 2019
- qFit-ligand reveals widespread conformational heterogeneity of drug-like molecules in X-ray electron density maps. G van Zundert, BM Hudson, S de Oliveira, DA Keedy, R Fonseca, A Heliou, P Suresh, K Borrelli, T Day, J Fraser, and H van den Bedem. J. Med. Chem. 2018
- Driving Structural Transitions in Molecular Simulations Using the Nonequilibrium Candidate Monte Carlo. A. Kurut, R. Fonseca, and W. Boomsma. J. Phys. Chem. B, 2018, 122 (3), pp 1195–1204
- Collision‐free poisson motion planning in ultra high‐dimensional molecular conformation spaces. R Fonseca, D Budday, H van den Bedem. Journal of Computational Chemistry. 2018. 10.1002/jcc.25138
- Frustration-guided motion planning reveals conformational transitions in proteins. D. Budday, R. Fonseca, S. Leyendecker and H. van den Bedem. Proteins. 2017, 85 1795-1807.
- Fast, clash-free RNA conformational morphing using molecular junctions. A. Heliou, D. Budday, R. Fonseca and H. van den Bedem. Bioinformatics. 2017, 33(14) 2114-2122.
- Probing RNA Native Conformational Ensembles with Structural Constraints. R. Fonseca, H. van den Bedem and J. Bernauer. Journal of Computational Biology. May 2016, 23(5): 362-371.
- Coupled Motions in β2AR:Gαs Conformational Ensembles. D.V. Pachov, R. Fonseca, D. Arnol, J. Bernauer, and H. van den Bedem. J. Chem. Theory Comput., 2016, 12 (3), pp 946-956.
- KGSrna: Efficient 3D kinematics-based sampling for nucleic acids. R. Fonseca, H. van den Bedem and J. Bernauer. Research in Computational Molecular Biology (LNCS) volume 9029, pages 80-95. 2015. [Supplementary material] [Video 1] [Video 2]
- Alpha Complexes in Protein Structure Prediction. P. Winter and R. Fonseca. Proc. of the International Conference on Bioinformatics Models, Methods and Algorithms. Pages 178-182. 2015
- Faster Exact Algorithms for Computing Steiner Trees in Higher Dimensional Euclidean Spaces. R. Fonseca, M. Brazil, P. Winter and M. Zachariasen. Proc. of the 11th DIMACS Implementation Challenge. 2014
- Steiner tree heuristics in Euclidean d-space. A. Olsen, S. Lorenzen, R. Fonseca and P. Winter. Proc. of the 11th DIMACS Implementation Challenge. 2014
- Characterizing RNA dynamics from NMR data with kinematic models. R. Fonseca, D.P. Pachov, J. Bernauer and H. van den Bedem. Nucleic Acids Research. 2014.
- Visualizing and representing the evolution of topological features. R. Fonseca and D. M. S. Jørgensen. Extended abstract for CG:YRF at Symposium of Computational Geometry. 2012. (pdf)
- Bounding Volumes for Proteins: A Comparative Study. R. Fonseca and P. Winter Journal of Computational Biology, 19(10): 1203-1213, 2012
- Ranking Beta Sheet Topologies with Applications to Protein Structure Prediction. R. Fonseca, G. Helles and P. Winter. Journal of Mathematical Modelling and Algorithms, 10(4): 357-369, 2011
- Protein Packing Quality using Delaunay Complexes. R. Fonseca. P. Winter and K. Karplus. Proceedings of ISVD 2011. Pages 117-122. 2011. (pdf)
- Adjustable Chain Trees for Proteins. P. Winter and R. Fonseca. Journal of Computational Biology. 19(1): 83-99. 2012
- Ranking Beta Sheet Topologies of Proteins. R. Fonseca, G. Helles and P. Winter. Proceedings of WCECS, 2:624-628. 2010.
- Protein Structure Prediction using Bee Colony Optimization Metaheuristic. R. Fonseca, M. Paluszewski and P. Winter. Journal of Molecular Modelling and Algorithms, 2010
- Predicting Dihedral Angles of Coil-structure Residues from the Primary Sequence using Neural Networks. G. Helles and R. Fonseca. BMC Bioinformatics 2009, 10:338
Links
- Solutions to small problems that I couldn't find implementations to anywhere else
- Java-implementation finding the cartesian coordinates of a regular n-simplex
- Java-implementation that solves the problem of Apollonius (i.e. finding a circle tangent to three other circles)
- Problem of Apollonius in 3D (i.e. finding a sphere tangent to four other spheres. Step-by-step description easily generalized to d-dimensions)
- Installing gnuplot (more importantly xz) on Mac with homebrew.
- Project ideas - Ideas I may never find the time to finish
- Lawsons paper for flip-based Delaunay triangulations - I was unable to google this paper so hopefully this will help someone else.